- java.lang.Object
-
- com.imsl.stat.Dissimilarities
-
- All Implemented Interfaces:
- Serializable, Cloneable
public class Dissimilarities extends Object implements Serializable, Cloneable
Computes a matrix of dissimilarities (or similarities) between the columns (or rows) of a matrix.Class
Dissimilaritiescomputes an upper triangular matrix (excluding the diagonal) of dissimilarities (or similarities) between the columns (or rows) of a matrix. Nine different distance measures can be computed. For the first three measures, three different scaling options can be employed. The distance matrix computed is generally used as input to clustering or multidimensional scaling functions.The following discussion assumes that the distance measure is being computed between the columns of the matrix. If distances between the rows of the matrix are desired, use
row=truein thesetRowmethod.The distance method and scaling option used by Dissimilarities can be set via methods
setDistanceMethodandsetScalingOption, respectively. For distance methodsL2_NORM,L1_NORM, orINFINITY_NORM, each row ofxis first scaled according to the value specified by thesetScalingOptionmethod. The scaling parameters are obtained from the values in the row scaled as either the standard deviation of the row or the row range; the standard deviation is computed from the unbiased estimate of the variance. If no scaling is performed, the parameters in the following discussion are all 1.0 (seesetScalingOption). Once the scaling value (if any) has been computed, the distance between column i and column j is computed via the difference vector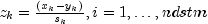 , where
, where 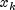 denotes the
k-th element in the i-th column,
denotes the
k-th element in the i-th column, 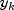 denotes the corresponding element in the j-th column, and ndstm
is the number of rows if differencing columns and the number of
columns if differencing rows. For given
denotes the corresponding element in the j-th column, and ndstm
is the number of rows if differencing columns and the number of
columns if differencing rows. For given 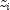 ,
the distance methods that allow scaling are defined as:
,
the distance methods that allow scaling are defined as:
The following distance measures do not allow for scaling.distanceMethodMetric L2_NORMEuclidean distance ( 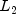 norm)
norm)L1_NORMSum of the absolute differences ( 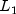 norm)
norm)INFINITY_NORMMaximum difference ( 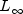 norm)
norm)distanceMethodMetric MAHALANOBISMahalanobis distance ABS_COSINEAbsolute value of the cosine of the angle between the vectors ANGLE_IN_RADIANSAngle in radians (0,  ) between the
lines through the origin defined by the vectors
) between the
lines through the origin defined by the vectorsCORRELATION_COEFFICIENTCorrelation coefficient ABS_CORRELATION_COEFFICIENTAbsolute value of the correlation coefficient EXACT_MATCHESNumber of exact matches, where 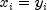 .
.For the Mahalanobis distance, any variable used in computing the distance measure that is (numerically) linearly dependent upon the previous variables in the
indexArrayvector from thesetIndexmethod is omitted from the distance measure.- See Also:
- Example 1, Example 2, Serialized Form
-
-
Nested Class Summary
Nested Classes Modifier and Type Class and Description static classDissimilarities.NoPositiveVarianceExceptionNo variable has positive variance.static classDissimilarities.ScaleFactorZeroExceptionThe computations cannot continue because a scale factor is zero.static classDissimilarities.ZeroNormExceptionThe computations cannot continue because the Euclidean norm of the column is equal to zero.
-
Field Summary
Fields Modifier and Type Field and Description static intABS_CORRELATION_COEFFICIENTIndicates the absolute value of the correlation coefficient distance method.static intABS_COSINEIndicates the absolute value of the cosine of the angle between the vectors distance method.static intANGLE_IN_RADIANSIndicates the angle in radians (0, ) between the lines through the
origin defined by the vectors distance method.
) between the lines through the
origin defined by the vectors distance method.static intCORRELATION_COEFFICIENTIndicates the correlation coefficient distance method.static intEXACT_MATCHESIndicates the number of exact matches distance method.static intINFINITY_NORMIndicates the maximum difference (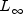 norm)
distance method.
norm)
distance method.static intL1_NORMIndicates the sum of the absolute differences (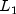 norm) distance method.
norm) distance method.static intL2_NORMIndicates the Euclidean distance method (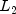 norm).
norm).static intMAHALANOBISIndicates the Mahalanobis distance method.static intNO_SCALINGIndicates no scaling.static intRANGEIndicates scaling by the range.static intSTD_DEVIndicates scaling by the standard deviation.
-
Constructor Summary
Constructors Constructor and Description Dissimilarities(double[][] x)Constructor forDissimilarities.
-
Method Summary
Methods Modifier and Type Method and Description voidcompute()Computes a matrix of dissimilarities (or similarities) between the columns (or rows) of a matrix.double[][]getDistanceMatrix()Returns the distance matrix.intgetDistanceMethod()Returns the method used in computing the dissimilarities or similarities.int[]getIndex()Returns the indices of the rows (columns) used in computing the distance measure.booleangetRow()Returns abooleanindicating whether distances are computed between rows or columns ofx.intgetScalingOption()Returns the scaling option.voidsetDistanceMethod(int distanceMethod)Sets the method to be used in computing the dissimilarities or similarities.voidsetIndex(int[] indexArray)Sets the indices of the rows (columns).voidsetRow(boolean row)Identifies whether distances are computed between rows or columns ofx.voidsetScalingOption(int distanceScale)Sets the scaling option used if theL2_NORM,L1_NORM, orINFINITY_NORMdistance methods are specified.
-
-
-
Field Detail
-
ABS_CORRELATION_COEFFICIENT
public static final int ABS_CORRELATION_COEFFICIENT
Indicates the absolute value of the correlation coefficient distance method.- See Also:
- Constant Field Values
-
ABS_COSINE
public static final int ABS_COSINE
Indicates the absolute value of the cosine of the angle between the vectors distance method.- See Also:
- Constant Field Values
-
ANGLE_IN_RADIANS
public static final int ANGLE_IN_RADIANS
Indicates the angle in radians (0, ) between the lines through the
origin defined by the vectors distance method.
) between the lines through the
origin defined by the vectors distance method.- See Also:
- Constant Field Values
-
CORRELATION_COEFFICIENT
public static final int CORRELATION_COEFFICIENT
Indicates the correlation coefficient distance method.- See Also:
- Constant Field Values
-
EXACT_MATCHES
public static final int EXACT_MATCHES
Indicates the number of exact matches distance method.- See Also:
- Constant Field Values
-
INFINITY_NORM
public static final int INFINITY_NORM
Indicates the maximum difference (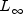 norm)
distance method.
norm)
distance method.- See Also:
- Constant Field Values
-
L1_NORM
public static final int L1_NORM
Indicates the sum of the absolute differences (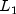 norm) distance method.
norm) distance method.- See Also:
- Constant Field Values
-
L2_NORM
public static final int L2_NORM
Indicates the Euclidean distance method (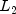 norm).
norm).- See Also:
- Constant Field Values
-
MAHALANOBIS
public static final int MAHALANOBIS
Indicates the Mahalanobis distance method.- See Also:
- Constant Field Values
-
NO_SCALING
public static final int NO_SCALING
Indicates no scaling.- See Also:
- Constant Field Values
-
RANGE
public static final int RANGE
Indicates scaling by the range.- See Also:
- Constant Field Values
-
STD_DEV
public static final int STD_DEV
Indicates scaling by the standard deviation.- See Also:
- Constant Field Values
-
-
Constructor Detail
-
Dissimilarities
public Dissimilarities(double[][] x)
Constructor forDissimilarities.- Parameters:
x- Adoublematrix containing the data input matrix.
-
-
Method Detail
-
compute
public void compute() throws Dissimilarities.ScaleFactorZeroException, Dissimilarities.ZeroNormException, Dissimilarities.NoPositiveVarianceExceptionComputes a matrix of dissimilarities (or similarities) between the columns (or rows) of a matrix.- Throws:
Dissimilarities.ScaleFactorZeroException- is thrown when computations cannot continue because a scale factor is zeroDissimilarities.NoPositiveVarianceException- is thrown when no variable has positive varianceDissimilarities.ZeroNormException- is thrown when the Euclidean norm of a column is equal to zero
-
getDistanceMatrix
public final double[][] getDistanceMatrix()
Returns the distance matrix.- Returns:
- A
doublematrix containing the distance matrix.
-
getDistanceMethod
public int getDistanceMethod()
Returns the method used in computing the dissimilarities or similarities.
-
getIndex
public int[] getIndex()
Returns the indices of the rows (columns) used in computing the distance measure.
-
getRow
public boolean getRow()
Returns abooleanindicating whether distances are computed between rows or columns ofx.
-
getScalingOption
public int getScalingOption()
Returns the scaling option.
-
setDistanceMethod
public void setDistanceMethod(int distanceMethod)
Sets the method to be used in computing the dissimilarities or similarities.- Parameters:
distanceMethod- Anintidentifying the method to be used in computing the dissimilarities or similarities. Acceptable values ofdistanceMethodare:
See class description for more details. By default,distanceMethodMetric L2_NORMEuclidean distance ( 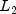 norm)
norm)L1_NORMSum of the absolute differences ( 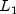 norm)
norm)INFINITY_NORMMaximum difference ( 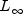 norm)
norm)MAHALANOBISMahalanobis distance ABS_COSINEAbsolute value of the cosine of the angle between the vectors ANGLE_IN_RADIANSAngle in radians (0,  ) between the
lines through the origin defined by the vectors
) between the
lines through the origin defined by the vectorsCORRELATION_COEFFICIENTCorrelation coefficient ABS_CORRELATION_COEFFICIENTAbsolute value of the correlation coefficient EXACT_MATCHESNumber of exact matches, where 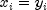 .
.distanceMethod=L2_NORM.
-
setIndex
public void setIndex(int[] indexArray)
Sets the indices of the rows (columns).- Parameters:
indexArray- Anintarray containing the indices of the columns (rows ifrow=false) to be used in computing the distance measure. By default, ifrow=true,indexArray=0, 1, ..., x[0].length-1. Ifrow=false,indexArray=0, 1, ..., x.length-1, seesetRow.
-
setRow
public void setRow(boolean row)
Identifies whether distances are computed between rows or columns ofx.- Parameters:
row- Abooleanidentifying whether distances are computed between rows or columns ofx. Ifrow=true, distances are computed between the rows ofx. Otherwise, distances between the columns ofxare computed. By default,row=true.
-
setScalingOption
public void setScalingOption(int distanceScale)
Sets the scaling option used if theL2_NORM,L1_NORM, orINFINITY_NORMdistance methods are specified. SeesetDistanceMethod.- Parameters:
distanceScale- Anintcontaining the scaling option. By default,distanceScale=NO_SCALING.distanceScaleMethod NO_SCALINGNo scaling is performed. STD_DEVIf setRow(false), scale each column by the standard deviation of the column.
IfsetRow(true), scale each row by the standard deviation of the row.RANGEIf setRow(false), scale each column by the range of the column.
IfsetRow(true), scale each row by the range of the row.
-
-