- java.lang.Object
-
- com.imsl.stat.ClusterKMeans
-
- All Implemented Interfaces:
- Serializable, Cloneable
public class ClusterKMeans extends Object implements Serializable, Cloneable
Perform a K-means (centroid) cluster analysis.ClusterKMeansis an implementation of Algorithm AS 136 by Hartigan and Wong (1979). It computes K-means (centroid) Euclidean metric clusters for an input matrix starting with initial estimates of the K cluster means. It allows for missing values (coded as NaN, not a number) and for weights and frequencies.Let p denote the number of variables to be used in computing the Euclidean distance between observations. The idea in K-means cluster analysis is to find a clustering (or grouping) of the observations so as to minimize the total within-cluster sums of squares. In this case, the total sums of squares within each cluster is computed as the sum of the centered sum of squares over all nonmissing values of each variable. That is,
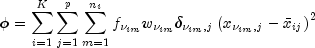
where
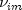 denotes the row index of the m-th observation
in the i-th cluster in the matrix X;
denotes the row index of the m-th observation
in the i-th cluster in the matrix X;
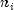 is the number of rows of X assigned to group i;
f denotes the frequency of the observation; w denotes its weight;
d is zero if the j-th variable on observation
is the number of rows of X assigned to group i;
f denotes the frequency of the observation; w denotes its weight;
d is zero if the j-th variable on observation 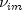 is missing, otherwise
is missing, otherwise 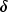 is one;
and
is one;
and 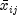 is the average of the
nonmissing observations for variable j in group i.
This method sequentially processes each observation and reassigns it to another
cluster if doing so results in a decrease in the total within-cluster sums of squares.
See Hartigan and Wong (1979) or Hartigan (1975) for details.
is the average of the
nonmissing observations for variable j in group i.
This method sequentially processes each observation and reassigns it to another
cluster if doing so results in a decrease in the total within-cluster sums of squares.
See Hartigan and Wong (1979) or Hartigan (1975) for details.- See Also:
- ClusterKMeans Example, Serialized Form
-
-
Nested Class Summary
Nested Classes Modifier and Type Class and Description static classClusterKMeans.ClusterNoPointsExceptionThere is a cluster with no pointsstatic classClusterKMeans.NoConvergenceExceptionConvergence did not occur within the maximum number of iterations.
-
Constructor Summary
Constructors Constructor and Description ClusterKMeans(double[][] x, double[][] cs)Constructor forClusterKMeans.
-
Method Summary
Methods Modifier and Type Method and Description double[][]compute()Computes the cluster means.int[]getClusterCounts()Returns the number of observations in each cluster.int[]getClusterMembership()Returns the cluster membership for each observation.double[]getClusterSSQ()Returns the within sum of squares for each cluster.voidsetFrequencies(double[] frequencies)Sets the frequency for each observation.voidsetMaxIterations(int iterations)Sets the maximum number of iterations.voidsetWeights(double[] weights)Sets the weight for each observation.
-
-
-
Constructor Detail
-
ClusterKMeans
public ClusterKMeans(double[][] x, double[][] cs)Constructor forClusterKMeans.- Parameters:
x- Adoublematrix containing the observations to be clustered.cs- Adoublematrix containing the cluster seeds, i.e. estimates for the cluster centers.
-
-
Method Detail
-
compute
public final double[][] compute() throws ClusterKMeans.NoConvergenceException, ClusterKMeans.ClusterNoPointsExceptionComputes the cluster means.- Returns:
- A
doublematrix containing computed result. - Throws:
com.imsl.stat.ClusterKMeans.NonnegativeFreqException- is thrown if a frequency is negative.com.imsl.stat.ClusterKMeans.NonnegativeWeightException- is thrown if a weight is negative.ClusterKMeans.NoConvergenceException- is thrown if convergence did not occur within the maximum number of iterations.ClusterKMeans.ClusterNoPointsException- is thrown if the cluster seed yields a cluster with no points.
-
getClusterCounts
public int[] getClusterCounts()
Returns the number of observations in each cluster. Note that thecomputemethod must be invoked first before invoking this method. Otherwise, the method throws aNullPointerExceptionexception.- Returns:
- An
intarray containing the number of observations in each cluster.
-
getClusterMembership
public int[] getClusterMembership()
Returns the cluster membership for each observation. Note that thecomputemethod must be invoked first before invoking this method. Otherwise, the method throws aNullPointerExceptionexception.- Returns:
- An
intarray containing the cluster membership for each observation. Cluster membership 1 indicates the observation belongs to cluster 1, cluster membership 2 indicates the observation belongs to cluster 2, etc.
-
getClusterSSQ
public double[] getClusterSSQ()
Returns the within sum of squares for each cluster. Note that thecomputemethod must be invoked first before invoking this method. Otherwise, the method throws aNullPointerExceptionexception.- Returns:
- A
doublearray containing the within sum of squares for each cluster.
-
setFrequencies
public void setFrequencies(double[] frequencies)
Sets the frequency for each observation.- Parameters:
frequencies- Adoublearray of sizex.lengthcontaining the frequency for each observation. Default:frequencies[]= 1.
-
setMaxIterations
public void setMaxIterations(int iterations)
Sets the maximum number of iterations.- Parameters:
iterations- Anintscalar specifying the maximum number of iterations. Default:interations= 30.
-
setWeights
public void setWeights(double[] weights)
Sets the weight for each observation.- Parameters:
weights- Adoublearray of sizex.lengthcontaining the weight for each observation. Default:weights[]= 1.
-
-