- java.lang.Object
-
- com.imsl.stat.ClusterHierarchical
-
- All Implemented Interfaces:
- Serializable, Cloneable
public class ClusterHierarchical extends Object implements Serializable, Cloneable
Performs a hierarchical cluster analysis from a distance matrix.Class
ClusterHierarchicalconducts a hierarchical cluster analysis based upon a distance matrix, or, by appropriate use of the transformation specified in the methodsetTransformType, based upon a similarity matrix. Only the upper triangular part of the input matrix is used.Hierarchical clustering in
ClusterHierarchicalproceeds as follows:
Initially, each data point is considered to be a cluster, numbered 1 ton, wherenis the number of rows in the input matrix,dist.- If the input matrix contains similarities, the matrix is transformed
to a distance matrix using the transform type specified by the method
setTransformType. Set k = 1. - A search is made of the distance matrix to find the two closest clusters. These clusters are merged to form a new cluster, numbered n + k. The cluster numbers of the two clusters joined at this stage are saved as Right Sons and Left Sons, and the distance measure between the two clusters is stored as Cluster Level.
- Based upon the method of clustering, updating of the distance
measure in the row and column of
distcorresponding to the new cluster is performed. - Set k = k + 1. If k is less than n, go to Step 2.
The five methods differ primarily in how the distance matrix is updated after two clusters have been joined. The argument
methodinsetMethodspecifies how the distance of the cluster just merged with each of the remaining clusters will be updated. ClassClusterHierarchicalallows five methods for computing the distances. To understand these measures, suppose in the following discussion that clusters A and B have just been joined to form cluster Z, and interest is in computing the distance of Z with another cluster called C.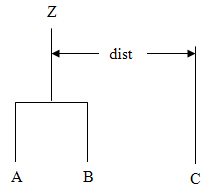
methodDescription LINKAGE_SINGLESingle linkage (minimum distance). The distance from Z to C is the minimum of the distances (A to C, B to C). LINKAGE_COMPLETEComplete linkage (maximum distance). The distance from Z to C is the maximum of the distances (A to C, B to C). LINKAGE_AVG_WITHIN_CLUSTERSAverage-distance-within-clusters method. The distance from Z to C is the average distance of all objects that would be within the cluster formed by merging clusters Z and C. This average may be computed according to formulas given by Anderberg (1973, page 139). LINKAGE_AVG_BETWEEN_CLUSTERSAverage-distance-between-clusters method. The distance from Z to C is the average distance of objects within cluster Z to objects within cluster C. This average may be computed according to methods given by Anderberg (1973, page 140). LINKAGE_WARDSWard's method: Clusters are formed so as to minimize the increase in the within-cluster sums of squares. The distance between two clusters is the increase in these sums of squares if the two clusters were merged. A method for computing this distance from a squared Euclidean distance matrix is given by Anderberg (1973, pages 142-145). In general, single linkage will yield long thin clusters while complete linkage will yield clusters that are more spherical. Average linkage and Ward's linkage tend to yield clusters that are similar to those obtained with complete linkage.
Class
ClusterHierarchicalproduces a unique representation of the binary cluster tree via the following three conventions; the fact that the tree is unique should aid in interpreting the clusters. First, when two clusters are joined and each cluster contains two or more data points, the cluster that was initially formed with the smallest level becomes the left son. Second, when a cluster containing more than one data point is joined with a cluster containing a single data point, the cluster with the single data point becomes the right son. Finally, when two clusters containing only one object are joined, the cluster with the smallest cluster number becomes the right son.Comments
- The clusters corresponding to the original data points are numbered
from 1 to
n, wherenis the number of rows indist. Then- 1 clusters formed by merging clusters are numberedn+ 1 ton+ (n- 1 ). - Raw correlations, if used as similarities, should be made positive
and transformed to a distance measure. One such transformation can
be performed by setting
transform=RECIPROCAL_ABS, in thesetTransformTypemethod. - The user may cluster either variables or observations with
ClusterHierarchicalsince a dissimilarity matrix, not the original data, is used. ClassDissimilaritiesmay be used to compute the matrixdistfor either the variables or observations.
- See Also:
- Example 1, Serialized Form
-
-
Field Summary
Fields Modifier and Type Field and Description static intLINKAGE_AVG_BETWEEN_CLUSTERSIndicates the average distance between (average distance between objects in the two clusters) method.static intLINKAGE_AVG_WITHIN_CLUSTERSIndicates the average distance within (average distance between objects within the merged cluster) method.static intLINKAGE_COMPLETEIndicates the complete linkage (maximum distance) method.static intLINKAGE_SINGLEIndicates the single linkage (minimum distance) method.static intLINKAGE_WARDSIndicates the Ward's method.static intMULTIPLICATIONIndicates transformation by multiplication by -1.0.static intNONEIndicates no transformation.static intRECIPROCAL_ABSIndicates transformation by taking the reciprocal of the absolute value.
-
Constructor Summary
Constructors Constructor and Description ClusterHierarchical(double[][] dist)Constructor forClusterHierarchical.
-
Method Summary
Methods Modifier and Type Method and Description voidcompute()Performs a hierarchical cluster analysis.int[]getClusterLeftSons()Returns the left sons of each merged cluster.double[]getClusterLevel()Returns the level at which the clusters are joined.int[]getClusterMembership(int nClusters)Returns the cluster membership of each observation.int[]getClusterRightSons()Returns the right sons of each merged cluster.intgetMethod()Returns the clustering method used.int[]getObsPerCluster(int nClusters)Returns the number of observations in each cluster.intgetTransformType()Returns the type of transformation.voidsetMethod(int method)Sets the clustering method to be used.voidsetTransformType(int transform)Sets the type of transformation.
-
-
-
Field Detail
-
LINKAGE_AVG_BETWEEN_CLUSTERS
public static final int LINKAGE_AVG_BETWEEN_CLUSTERS
Indicates the average distance between (average distance between objects in the two clusters) method.- See Also:
- Constant Field Values
-
LINKAGE_AVG_WITHIN_CLUSTERS
public static final int LINKAGE_AVG_WITHIN_CLUSTERS
Indicates the average distance within (average distance between objects within the merged cluster) method.- See Also:
- Constant Field Values
-
LINKAGE_COMPLETE
public static final int LINKAGE_COMPLETE
Indicates the complete linkage (maximum distance) method.- See Also:
- Constant Field Values
-
LINKAGE_SINGLE
public static final int LINKAGE_SINGLE
Indicates the single linkage (minimum distance) method.- See Also:
- Constant Field Values
-
LINKAGE_WARDS
public static final int LINKAGE_WARDS
Indicates the Ward's method.- See Also:
- Constant Field Values
-
MULTIPLICATION
public static final int MULTIPLICATION
Indicates transformation by multiplication by -1.0.- See Also:
- Constant Field Values
-
NONE
public static final int NONE
Indicates no transformation.- See Also:
- Constant Field Values
-
RECIPROCAL_ABS
public static final int RECIPROCAL_ABS
Indicates transformation by taking the reciprocal of the absolute value.- See Also:
- Constant Field Values
-
-
Constructor Detail
-
ClusterHierarchical
public ClusterHierarchical(double[][] dist)
Constructor forClusterHierarchical.- Parameters:
dist- Adoublesymmetric matrix containing the distance (or similarity) matrix. Only the upper triangular part is used.
-
-
Method Detail
-
compute
public void compute()
Performs a hierarchical cluster analysis.
-
getClusterLeftSons
public final int[] getClusterLeftSons()
Returns the left sons of each merged cluster.- Returns:
- An
intarray containing the left sons of each merged cluster.
-
getClusterLevel
public final double[] getClusterLevel()
Returns the level at which the clusters are joined.- Returns:
- A
doublearray containing the level at which the clusters are joined. Element [k-1] contains the distance (or similarity) level at which clustern+ k was formed.
-
getClusterMembership
public final int[] getClusterMembership(int nClusters)
Returns the cluster membership of each observation.- Parameters:
nClusters- Anintwhich specifies the desired number of clusters.- Returns:
- An
intarray containing the cluster membership of each observation.
-
getClusterRightSons
public final int[] getClusterRightSons()
Returns the right sons of each merged cluster.- Returns:
- An
intarray containing the right sons of each merged cluster.
-
getMethod
public int getMethod()
Returns the clustering method used.
-
getObsPerCluster
public final int[] getObsPerCluster(int nClusters)
Returns the number of observations in each cluster.- Parameters:
nClusters- Anintwhich specifies the desired number of clusters.- Returns:
- An
intarray containing the number of observations in each cluster.
-
getTransformType
public int getTransformType()
Returns the type of transformation.
-
setMethod
public void setMethod(int method)
Sets the clustering method to be used.- Parameters:
method- Anintidentifying the clustering method to be used. By default,method=LINKAGE_SINGLE.methodDescription LINKAGE_SINGLESingle linkage (minimum distance). LINKAGE_COMPLETEComplete linkage (maximum distance). LINKAGE_AVG_WITHIN_CLUSTERSAverage distance within (average distance between objects within the merged cluster). LINKAGE_AVG_BETWEEN_CLUSTERSAverage distance between (average distance between objects in the two clusters). LINKAGE_WARDSWard's method (minimize the within-cluster sums of squares). For Ward's method, the elements of distare assumed to be Euclidean distances.
-
setTransformType
public void setTransformType(int transform)
Sets the type of transformation.- Parameters:
transform- Anintidentifying the type of transformation applied to the measures indist. By default,transform=NONE.transformDescription NONENo transformation is required. The elements of distare distances.MULTIPLICATIONConvert similarities to distances by multiplication by -1.0. RECIPROCAL_ABSConvert similarities (usually correlations) to distances by taking the reciprocal of the absolute value.
-
-